In a previous post I gave my recommendation to use signed rather unsigned networks. This post will describe the two slightly different formulas that WGCNA offers for building signed networks from a correlation matrix. As a quick reminder, constructing a network really means calculating its adjacency matrix aij. Elements of this matrix encode the connection strengths between nodes of the network and must lie between 0 (unconnected) and 1 (fully connected). The “signed” network adjacency 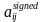 scales the correlation to lie between 0 and 1, then raises it to a power:
scales the correlation to lie between 0 and 1, then raises it to a power:
![]()
A salient feature of the signed adjacency is that zero correlation gives rise to a (theoretically) non-zero adjacency (connection strength). Although at first sight counter-intuitive, this can sometimes be desirable, for example in sample networks. However, in most applications to large, high-throughput data, the power β is sufficiently large (usually at least 10-12) that negative and small positive correlations lead to negligibly small adjacencies. The figure below illustrates this point.
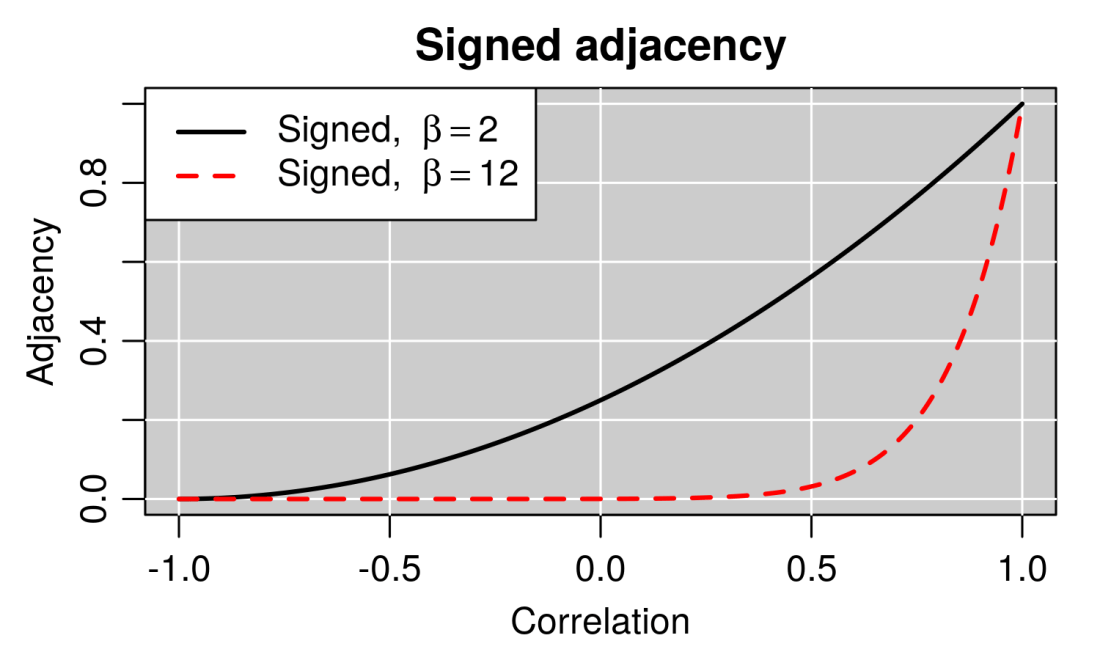 |
| Signed adjacency (y-axis) as a function of correlation (x-axis) for two soft thresholding powers β. When the power is sufficiently high (say 12, red line), the adjacency is negligibly small for all negative and small positive correlations. |
Unlike the “signed” adjacency, the “signed hybrid” adjacency is exactly zero for all negative (or zero) correlations:

We call this adjacency “hybrid” because it uses a combination (hybrid) of hard and soft thresholding: there is a hard threshold at 0, and soft thresholding above zero. The figure below illustrates the signed hybrid adjacency for soft thresholding powers 1 and 6.
 |
| Signed hybrid adjacency (y-axis) as a function of correlation (x-axis). This adjacency function leads to zero adjacencies for all negative correlations and all choices of β. Choosing β=6 also suppresses low positive correlations. |
And which one should you use? In high throughput data, with more variables (genes) than observation (samples), one would typically use a fairly high soft thresholding power β, say β=12 for a signed or β=6 for a signed hybrid network. While the formulas look quite different, the results for these two powers will be extremely similar:
 |
| Solid red line shows signed adjacency obtained with power β=12, dashed cyan line shows signed hybrid adjacency obtained with power β=6. The two adjacencies are nearly indistinguishable. |
Thus, in the end the two signed network variants result in nearly identical networks as long as (1) the signed network uses twice the soft thresholding power of the signed hybrid network, and (2) the power is suitable for analysis of high-throughput, more-variables-than-samples, data: at least 4 for signed hybrid, and at least 8 for signed networks. In other words, use either but remember to double the power for the signed, compared to the signed hybrid.